This function produces a boxplot of the residuals from WRTDS, expressed in natural log concentration units. It provides an alternative for viewing the standardized residuals, where the each residual is divided by its estimated standard error. The monthly boxplot widths are proportional to the square root of the sample size. The residuals for a censored value are determined as the difference between the natural log of the average of the upper and lower. bounds on the sample value, minus the log space estimate of concentration.
Although there are a lot of optional arguments to this function, most are set to a logical default.
Data come from named list, which contains a Sample dataframe with the sample data, and an INFO dataframe with metadata
Usage
boxResidMonth(eList, stdResid = FALSE, las = 1, printTitle = TRUE,
cex = 0.8, cex.axis = 1.1, cex.main = 1.1, font.main = 2,
tinyPlot = FALSE, customPar = FALSE, monthLab = 1,
randomCensored = FALSE, ...)Arguments
- eList
named list with at least the Sample and INFO dataframes
- stdResid
logical variable, if TRUE it uses the standardized residual, if FALSE it uses the actual, default is FALSE
- las
numeric in c(0,1,2,3); the style of axis labels
- printTitle
logical variable if TRUE title is printed, if FALSE not printed (this is best for a multi-plot figure)
- cex
numerical value giving the amount by which plotting symbols should be magnified
- cex.axis
magnification to be used for axis annotation relative to the current setting of cex
- cex.main
magnification to be used for main titles relative to the current setting of cex
- font.main
font to be used for plot main titles
- tinyPlot
logical variable, if TRUE plot is designed to be plotted small, as a part of a multipart figure, default is FALSE
- customPar
logical defaults to FALSE. If TRUE, par() should be set by user before calling this function
- monthLab
object of monthLabel class, or numeric represented the short code, or character representing the descriptive name.
- randomCensored
logical. Show censored residuals as randomized. Default = FALSE.
- ...
arbitrary graphical parameters that will be passed to genericEGRETDotPlot function (see ?par for options)
Examples
eList <- Choptank_eList
# Water year:
boxResidMonth(eList)
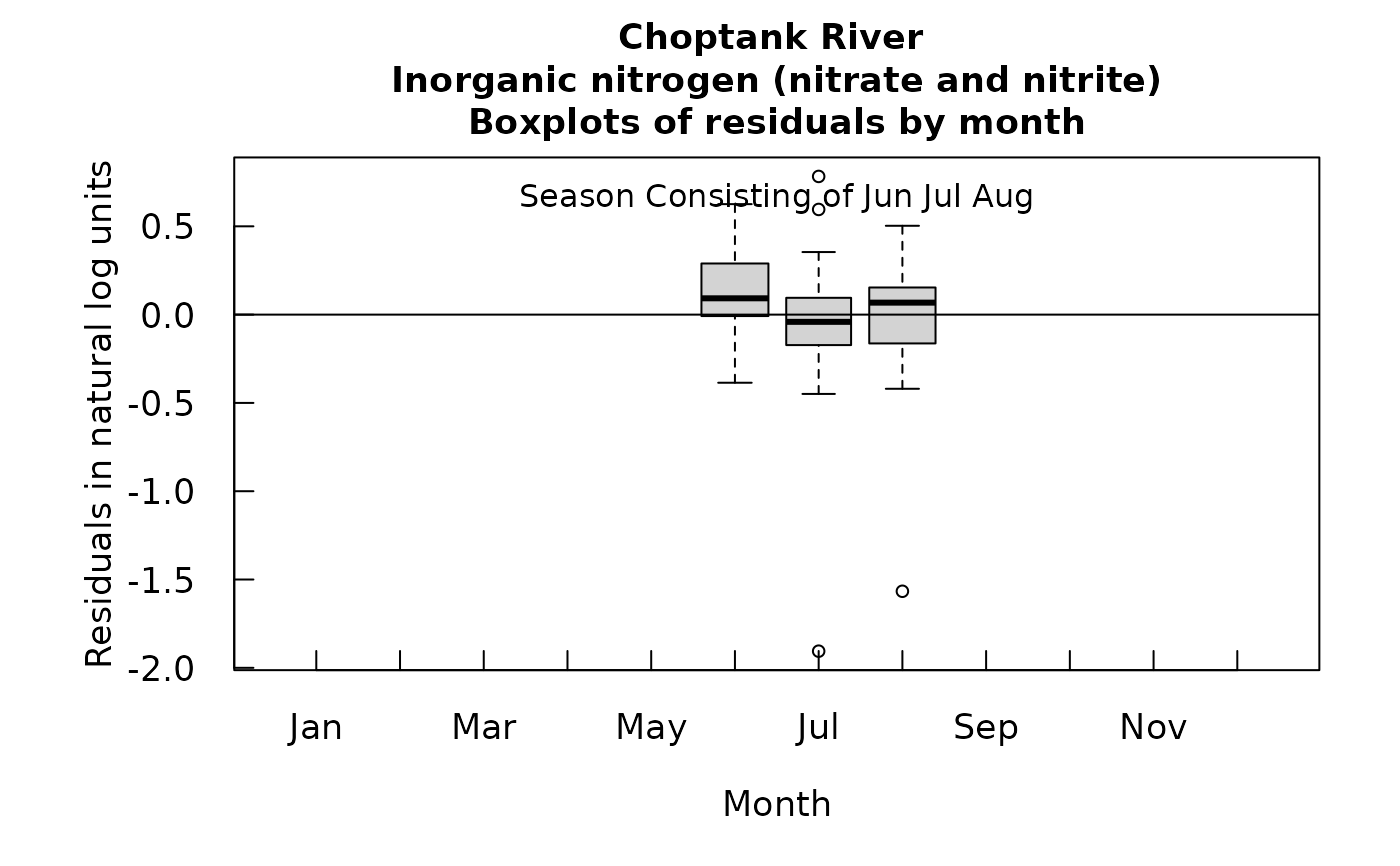
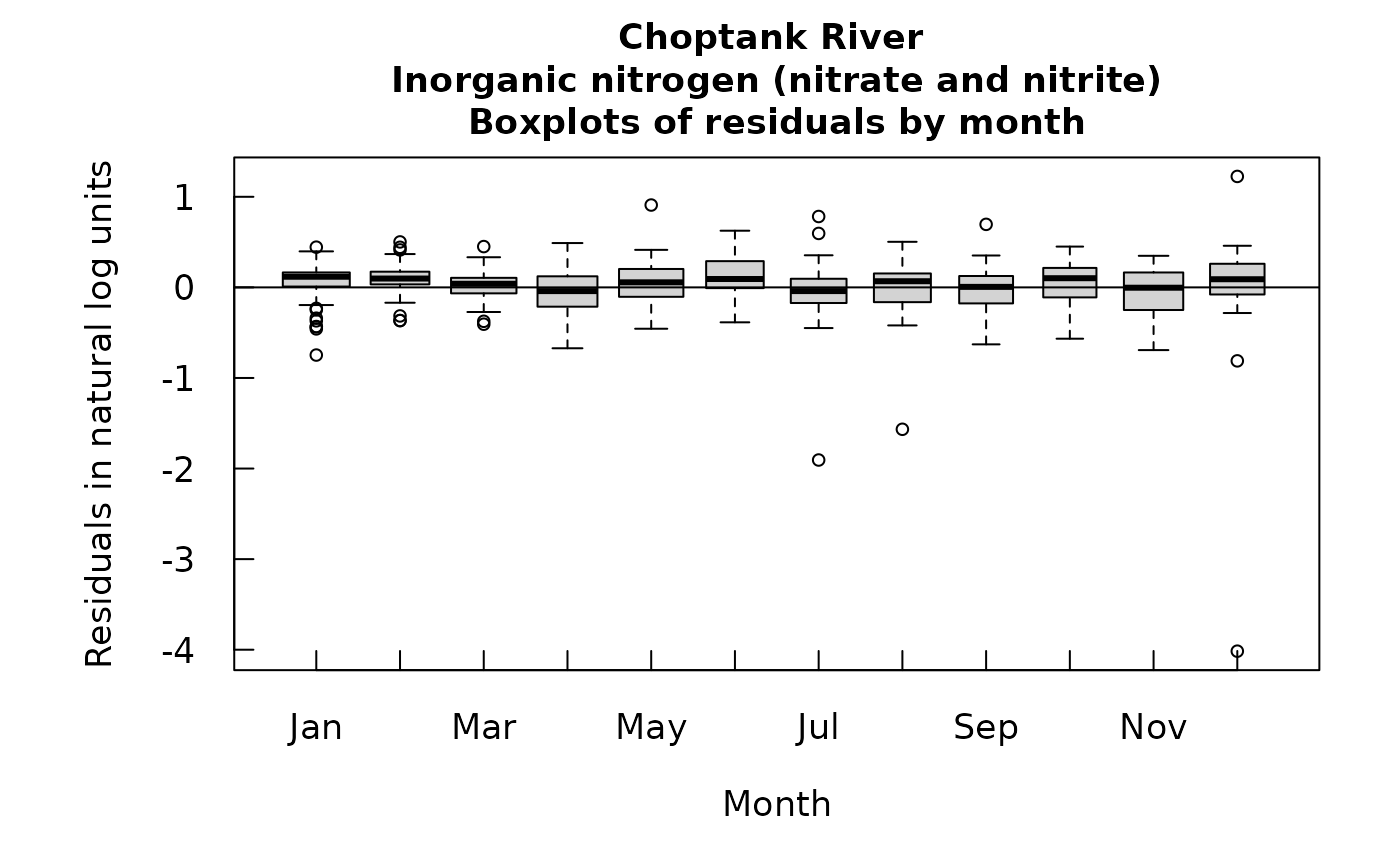 # Graphs consisting of Jun-Aug
eList <- setPA(eList, paStart = 6, paLong = 3)
boxResidMonth(eList)
# Graphs consisting of Jun-Aug
eList <- setPA(eList, paStart = 6, paLong = 3)
boxResidMonth(eList)

